Researchers discover biosynthetic gene of archaeal membrane lipid GMGT
DATE:2024-06-21
Most archaeal cell membranes are composed of tetraether lipids known as GDGTs (Glycerol Dialkyl Glycerol Tetraethers). GDGTs form a monolayer cell membrane that exhibits excellent physicochemical stability, significantly enhancing the ability of archaea to adapt to extreme environments. Additionally, GDGTs are important biomarkers widely used in the study of paleoclimate changes.
GDGTs include various derivatives, such as cyclized, hydroxylated, and methylated forms, among which one significant derivative is the H-shape GDGT, also known as GMGT (Glycerol Monoalkyl Glycerol Tetraether). However, the biosynthetic pathways, physiological functions, and potential geoscience significance of GMGTs remain unknown.
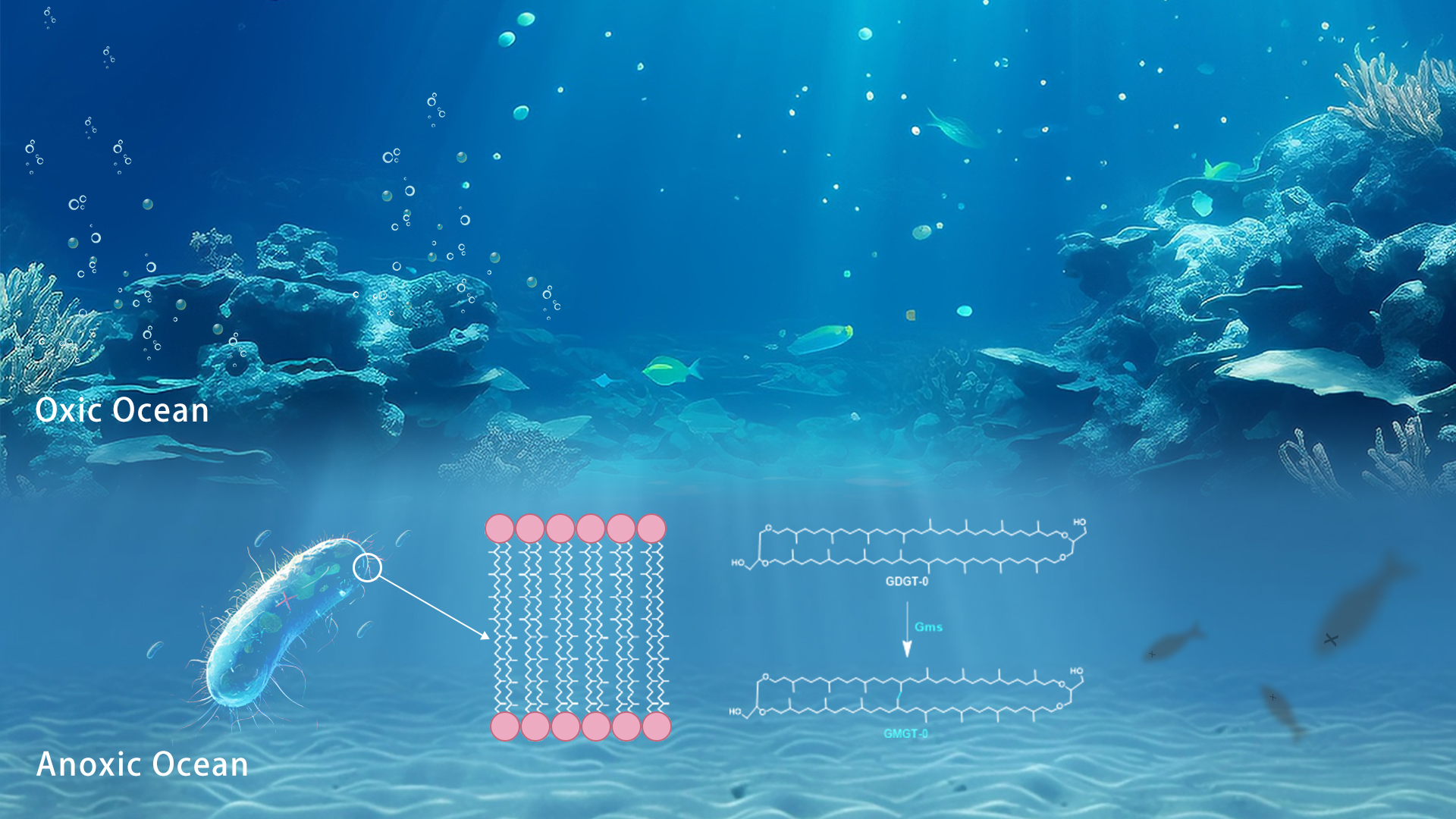

A collaborative research team led by Associate Professor Zhirui Zeng from the Department of Ocean Science and Engineering (OSE), Associate Professor Lizhi Tao from the Department of Chemistry, and Assistant Professor Fuxing Zeng from the Department of Systems Biology, all of the Southern University of Science and Technology (SUSTech), have discovered the biosynthetic gene of archaeal membrane lipid GMGT.
Their work, entitled “Biosynthesis of GMGT lipids by a radical SAM enzyme associated with anaerobic archaea and oxygen-deficient environments”, has been published in Nature Communications, a multidisciplinary journal that covers the natural sciences, including physics, chemistry, earth sciences, medicine, and biology.
In this study, the researchers combined in vivo heterologous gene expression in archaea, and in vitro biochemical activity assays to identify, for the first time, the protein responsible for the GMGT biosynthesis (GMGT synthase, Gms) (Figure 1). Gms belongs to the radical SAM superfamily, utilizing free radicals to activate the C(sp3)-H bond of the GDGT substrate and constructing a C(sp3)-C(sp3) bond to form the product GMGT.
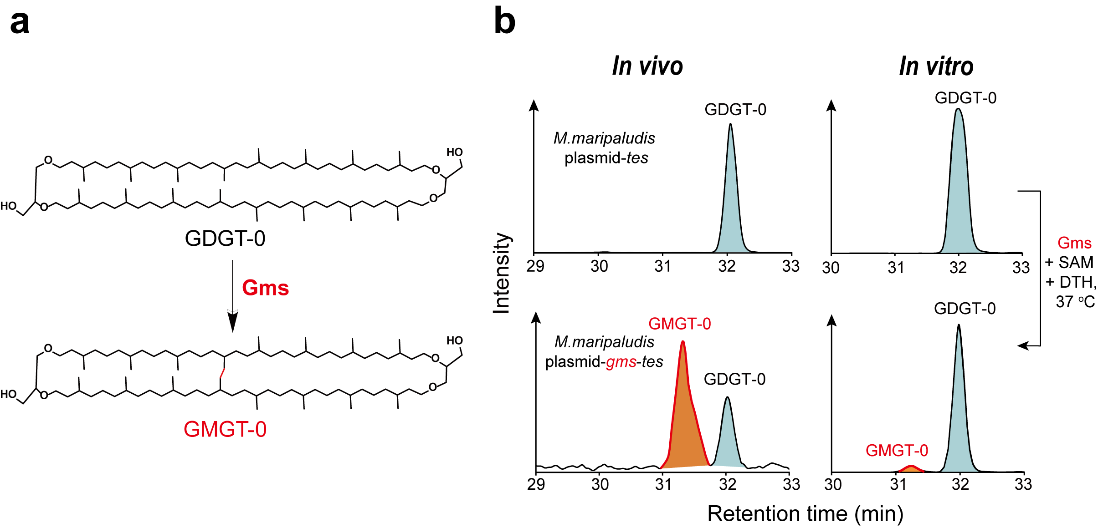
Figure 1. The in vivo and in vitro assays identify Gms as responsible for GMGT biosynthesis
To explore the catalytic details of the Gms enzyme, they used electron paramagnetic resonance (EPR) spectroscopy analysis to confirm that the three FeS clusters in Gms are crucial for electron transfer and the construction of the C(sp3)-C(sp3) bond. Alphafold2 structure prediction revealed the unique substrate-binding space of Gms and discovered that the cap structure formed by the N-terminal sequence may play an important role in stabilizing the substrate binding (Figure 2).
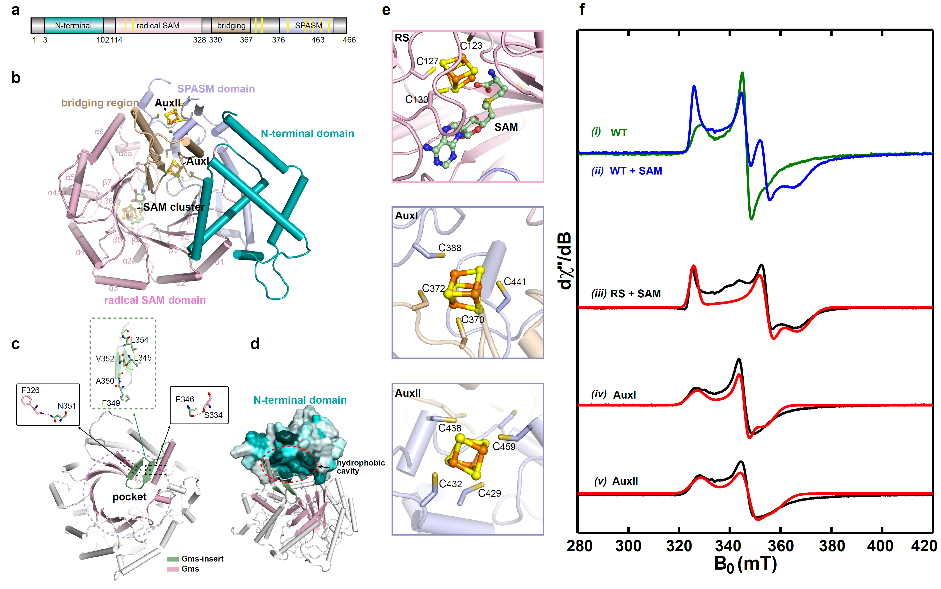
Figure 2. Alphafold2 structure prediction and EPR analysis of Gms
The discovery of GMGT synthase enables the identification of the biological source of GMGT. Through Gms homologous search in genomes (BLASTP), they found that the strains synthesizing GMGT are all restricted anaerobic archaea. Additionally, Gms gene is exclusively present in the metagenomes from oxygen-deficient environments. These findings suggest Gms has the potential to record environmental, especially marine, anoxia history (Figure 3).
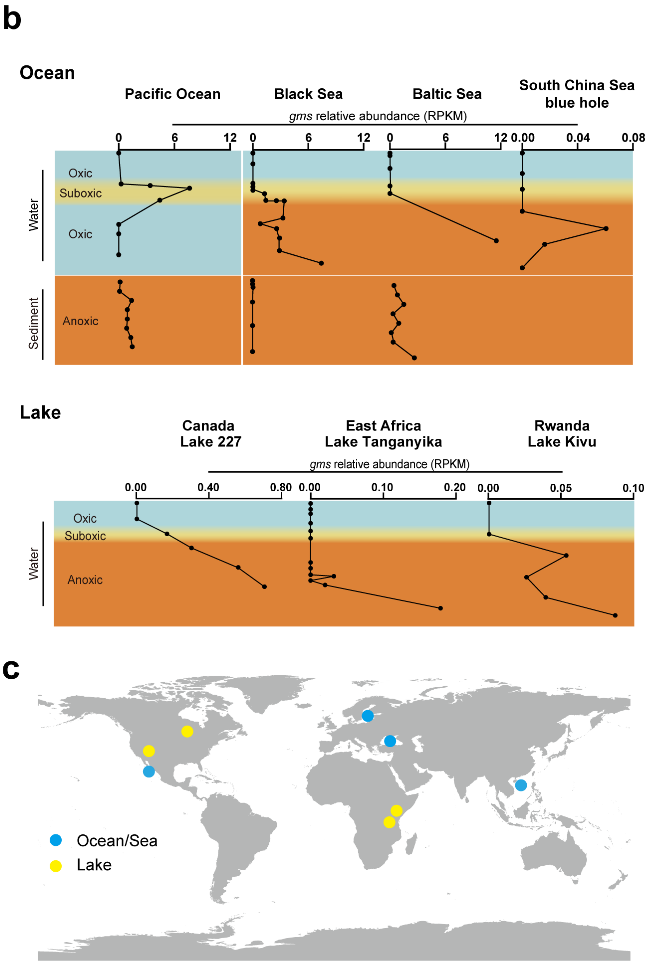
Figure 3. Gms gene is exclusively present in oxygen-deficient environments
Former Research Assistant Yanan Li from the Department of OSE at SUSTech is the first author of the paper. Associate Professors Zhirui Zeng and Lizhi Tao, along with Assistant Professor Fuxing Zeng, are the co-corresponding authors. Co-authors include Ting Yu from the Department of Systems Biology at SUSTech, Xi Feng, Bo Zhao, and Huahui Chen from the Department of OSE at SUSTech, Associate Professor Huan Yang from China University of Geosciences (Wuhan), Professor Xiao-Hua Zhang and doctoral student Xing Chen from the Ocean University of China, and Associate Professor Noah Burns and doctoral student Hayden Anderson from Stanford University.
The study was supported by the National Natural Science Foundation of China (NSFC).
Paper link: https://doi.org/10.1038/s41467-024-49650-x
To read all stories about SUSTech science, subscribe to the monthly SUSTech Newsletter.
latest news
-
SUSTech team identifies transposable elements as key determinants of 3D genome structure
Date:2026-03-16
-
SUSTech Researchers Uncover the Molecular Mechanism of C1ql1/BAI3 Assemblies in CF-PC Synapse Development
Date:2025-12-29
-
Researchers discover mechanisms underlying how gravity competes with vision to guide brain’s inner compass
Date:2025-09-22
-
Dynamic changes in transposable elements shape human three-germ-layer differentiation
Date:2025-09-04
